Structure-based identification of SARS-derived peptides with potential to induce broad protective immunity
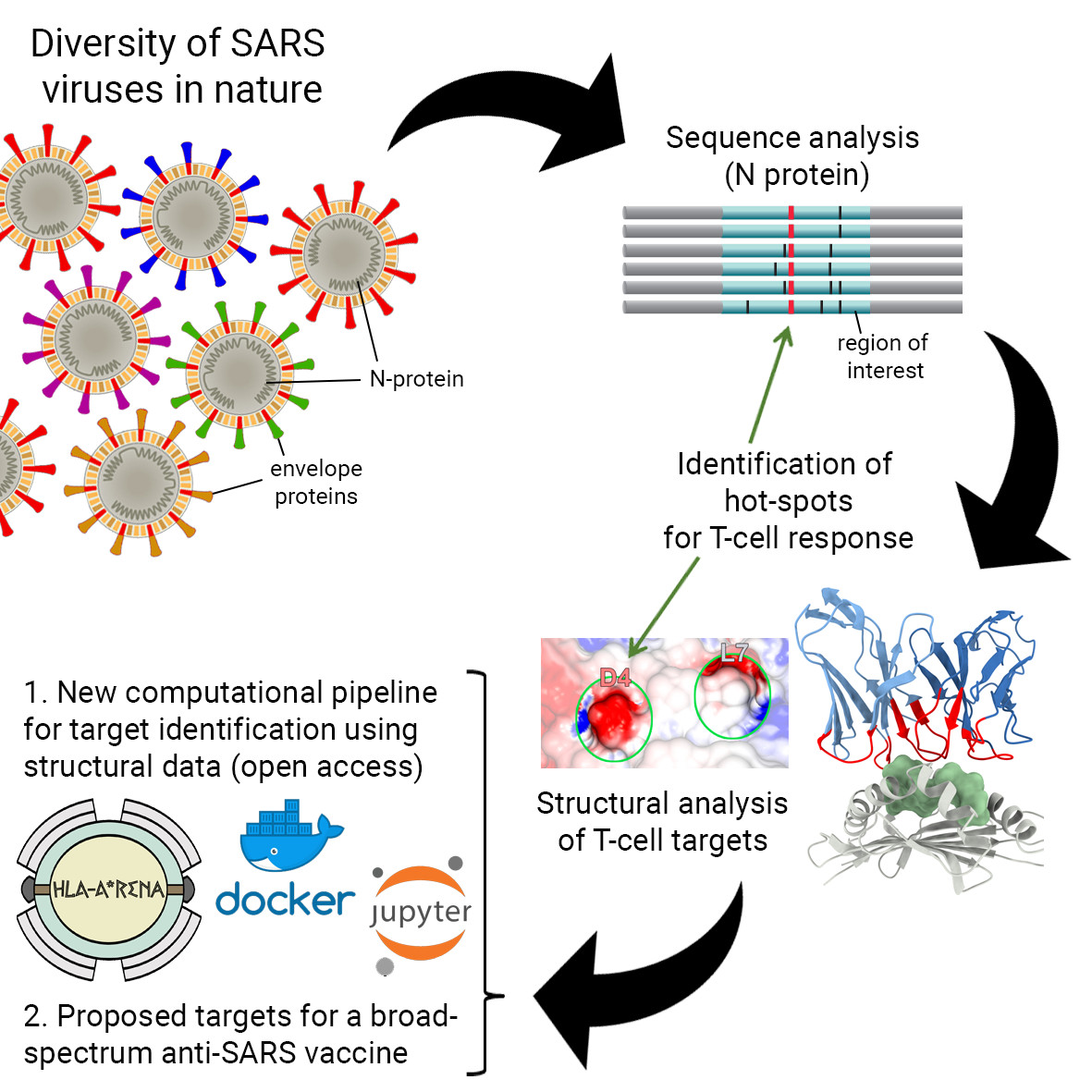
In 2019, a pandemic age caused by a novel strain of coronavirus (SARS-CoV-2) with a fast-growing number of confirmed cases all over the world took place. Several efforts are underway to produce a new vaccine, promoting immediate and long-term cell-mediated immunity based on T cell lymphocytes. T cell responses are particularly important for fighting viral infections because they can find and eliminate infected cells. This project will develop a computational pipeline to enable the identification of peptides that are conserved across different SARS-CoV strains, and that can potentially be used to induce broad protective cellular immunity against these viruses. The approach is based on the combined use of gold-standard sequence-based methods, and new cutting-edge methods for the structural modeling and analysis of peptides bound to different Human Leukocyte Antigen (HLA) receptors. HLAs are responsible for displaying the peptides to T-cell lymphocytes, and the proposed pipeline will enable the identification of conserved hot-spots capable of triggering T-cell responses against multiple SARS-CoV variants. In the context of this project, research will target conserved peptides from the Nucleocapsid (N) protein of SARS-CoV-2. If needed, the optimization of predicted peptides will be conducted for different prevalent HLA alleles. The proposed computational pipeline will be built using general software-engineering principles, making it also applicable to study different proteins from SARS-CoV variants, and even other pathogens. For more information you can access NSF Covid Information Commons webpage at https://covidinfocommons.datascience.columbia.edu/awards/2033262.
This work has been supported by grant NSF DBI 2033262.
Related Publications
- M. M. Rigo, R. Fasoulis, A. Conev, S. Hall-Swan, D. Amaral Antunes, and L. Kavraki, “SARS-Arena: Sequence and Structure-Guided Selection of Conserved Peptides from
SARS-related Coronaviruses for Novel Vaccine Development,” Frontiers in Immunology, vol. 13, Jul. 2022.
Details - R. F. Tarabini, M. M. Rigo, A. Faustino Fonseca, F. Rubin, R. Bellé, L. E. Kavraki, T. C. Ferreto, D. Amaral Antunes, and A. P. D. de Souza, “Large-Scale Structure-Based Screening of Potential T Cell Cross-Reactivities Involving
Peptide-Targets From BCG Vaccine and SARS-CoV-2,” Frontiers in Immunology, vol. 12, Jan. 2022.
Details - S. Hall-Swan, D. Devaurs, M. M. Rigo, D. A. Antunes, L. E. Kavraki, and G. Zanatta, “DINC-COVID: A webserver for ensemble docking with flexible SARS-CoV-2 proteins,” Computers in Biology and Medicine, vol. 139, p. 104943, 2021.
Details